All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit the International Myeloma Foundation or HealthTree for Multiple Myeloma.
The Multiple Myeloma Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the Multiple Myeloma Hub cannot guarantee the accuracy of translated content. The Multiple Myeloma Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The Multiple Myeloma Hub is an independent medical education platform, sponsored by Bristol Myers Squibb, GSK, Legend Biotech, Pfizer, and Roche. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View multiple myeloma content recommended for you
Risk stratification of newly diagnosed multiple myeloma using chromosome 1
Do you know... According to a study by Yang et al.2, which cytogenetic abnormality when present in patients with NDMM with gain of chromosome 1q negatively impacts overall survival but not progression-free survival?
Abnormalities of chromosome 1 are common genetic mutations in patients with multiple myeloma (MM).1 While some studies have reported worse outcomes for patients with chromosome 1 anomalies, prevalence, real-world treatment, and outcomes are unknown.1
With the possibility of chromosome 1 abnormalities being adopted into future International Staging System criteria for newly diagnosed MM,1 the Multiple Myeloma Hub explores recent current evidence on its use as a prognostic marker.
The hallmark of MM is abnormal clonal plasma cells, whose biology and behavior have been affected by sequential genetic changes, leading to disease onset, progression, and treatment resistance.1 The genomic and epigenomic profiles seen in newly diagnosed MM (NDMM) are highly heterogeneous, with further complex genomic changes arising from chromosomal instability (CIN), microsatellite instability, and increasing numbers of gene mutations.1,2 The Multiple Myeloma Hub has previously covered chromosome 1 abnormalities in NDMM, alternative staging systems (e.g., the Mayo Additive Scoring System) and incorporating them, and the need to consider their relevance to risk and prognosis.
CIN changes in chromosome 1 are the most frequent copy number variants (CNV) and structural changes seen in MM, with 1q gain the most common cytogenetic abnormality.1,2 Anomalies involving a whole arm of 1q and 1p deletion are associated with high-risk, poor prognosis MM, but with variable and inconsistent significance.1,2 The frequent occurrence of 1q anomalies alongside other high-risk cytogenetic abnormalities, for example del(17p), has raised questions over whether it is a driver of inferior outcomes or occurs alongside them as a result of the genomic instability considered above.1,2
To address this topic, the Multiple Myeloma Hub is pleased to summarize the three following publications:
- A review by Luo et al.1, published in the Federation of American Societies for Experimental Biology Journal in April 2022, which describes the role of chromosome 1 instability in the progression of MM.
- A study by Yang et al.2, published in the American Journal of Hematology in August 2022, which explored 1q gain as an independent risk factor for scoring patients with MM.
- A study by Kastritis et al.3, which considered chromosome 1q21 aberrations in identifying ultra-high-risk disease, published in the American Journal of Hematology in May 2022.
Chromosome 1 in multiple myeloma biology1
Chromosomal instability can lead to structural chromosomal changes and CNV, genetic events which can be considered as primary or secondary. Those present in early phases of MM (e.g., monoclonal gammopathy of unknown significance) are considered primary, while those identified only later in the disease course are secondary. Primary genomic events commonly include hyperdiploidy or translocations, while secondary events include chromosomal translocations, CNV, and single-nucleotide variants.
CNV result in the loss and gain of DNA, at the level of either an entire chromosome arm or a small region of amplification or deletion; while isochromosome anomalies see the duplication of one arm and loss of the corresponding chromosome arm. Both the short and long arms of chromosome 1 can be involved, especially 1q gain (35–40% of patients) and 1p deletions, affecting disease biology and clinical outcomes (Figure 1).
Figure 1. Effects of chromosome 1 instability on clinical outcomes in patients with NDMM*
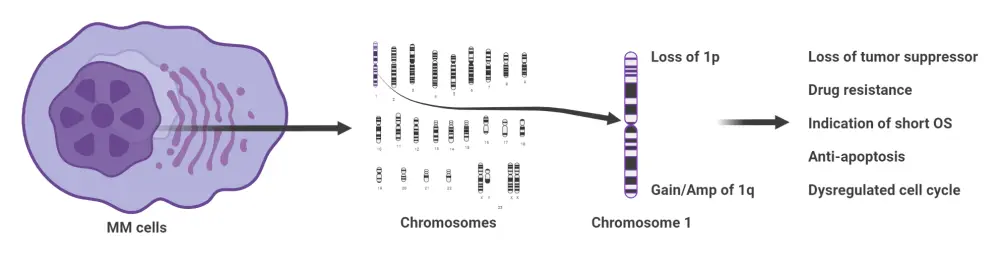
Amp, amplification; MM, multiple myeloma; NDMM, newly diagnosed multiple myeloma; OS, overall survival.
*Adapted from Luo, et al.1 Created with BioRender.com.
Changes in DNA structure or integrity arising from CIN manifest as changes in gene behavior and expression, giving rise to the disease process of MM and the behavior of cell lineages and clones as the disease progresses. Central to understanding the relevance of chromosome 1 anomalies to different elements of MM biology is a consideration of the genes affected, some of which are listed in Table 1. Some 1p losses are associated with loss of tumor suppressor function, while upregulation from 1q gain can result in upregulation. Some affected genes even hold the potential to be therapeutic targets (e.g., CD46).
Table 1. Genes affected by chromosome 1 instability in multiple myeloma*
|
MM, multiple myeloma; mRNA, messenger RNA. |
||||
|
Region |
Locus |
Gene |
Biological function |
Role in MM |
|---|---|---|---|---|
|
Chromosome 1p12 |
1p12 |
FAM464C |
Enhances mRNA stability and gene expression |
Tumor suppressor |
|
Chromosome 1p31 |
1p31.3 |
USP33 |
Regulates centrosome homeostasis |
— |
|
Chromosome 1p32 |
1p32.3 |
CDKN2C |
A D-type cyclin-dependent kinase inhibitor, involved in plasma cell homeostasis |
Cell cycle regulation |
|
Chromosome 1p36 |
1p36.32 |
TP73 |
Participates in the apoptotic response to DNA damage |
Promotion of apoptosis |
|
Chromosome 1q12-1q23 |
1q21.1 |
PDZK1 |
Organizing proteins at the cell membrane, linking transmembrane proteins to the actin cytoskeleton |
Overexpression contributes to malignant phenotype including drug resistance |
|
Chromosome 1q12-1q23 |
1q21.3 |
ADAR1 |
Catalyzes the hydrolytic deamination of adenosine to inosine in double-stranded RNA |
Oncogenic, driving cellular growth and proliferation in MM |
|
Chromosome 1q31-1qter |
1q32.3 |
CD46 |
Regulates the adaptive immune response |
Potential surface marker for targeted cytoreductive therapy of MM |
Chromosome 1q gain in multiple myeloma2
To explore the clinical significance of 1q gain in NDMM, Yang et al.2 performed a retrospective analysis of 934 patients with NDMM treated in two hospitals between November 2009 and November 2019. All patients received proteosome inhibitors or immunomodulatory drugs in first-line treatment. The study design included a planned replication study, using demographic and clinical data from the CoMMpass trial (NCT01454297).
With a median follow-up of 44.1 months, chromosome 1q gain was identified in 53.1% of patients. Patient data can be seen in Table 2.
Table 2. Clinical and demographic differences between patients with and without chromosome 1q*
|
β2-MG, serum β2-microglobulin; BMPC, bone marrow plasma cell; del, deletion; Hb, hemoglobin; ISS, International Staging System; LDH, lactate dehydrogenase; M protein, monoclonal protein type; R-ISS, revised International Staging System; t, translocation. |
|||
|
Characteristic, % (unless stated otherwise) |
Absence of 1q gain |
Presence of 1q gain |
p value |
|---|---|---|---|
|
Median age (range), years |
61 (27–87) |
60 (32–84) |
|
|
Age ≥65 years |
35.4 |
31.5 |
|
|
Sex |
0.207 |
||
|
Male |
60.7 |
56.7 |
|
|
Female |
39.3 |
43.3 |
|
|
M protein |
<0.001 |
||
|
IgG |
48.6 |
41.9 |
0.040 |
|
IgA |
20.3 |
27.8 |
0.008 |
|
IgD |
3.2 |
9.5 |
<0.001 |
|
ISS |
0.013 |
||
|
I |
18.7 |
12.3 |
0.007 |
|
II |
31.7 |
30.8 |
0.770 |
|
III |
49.5 |
56.9 |
0.025 |
|
R-ISS |
0.004 |
||
|
I |
12.3 |
7.7 |
0.025 |
|
II |
63.4 |
58.8 |
0.180 |
|
III |
24.3 |
33.6 |
0.004 |
|
BMPCs |
0.008 |
||
|
≥30 |
60.2 |
71.3 |
|
|
<30 |
39.8 |
28.7 |
|
|
β2-MG, mg/L |
0.003 |
||
|
≥5.5 |
50.6 |
63.6 |
|
|
<5.5 |
49.4 |
36.4 |
|
|
LDH, U/L |
<0.001 |
||
|
≥220 |
22.1 |
33.1 |
|
|
<220 |
77.9 |
66.9 |
|
|
Hb, g/L |
<0.001 |
||
|
<100 |
62.6 |
74.8 |
|
|
≥100 |
37.4 |
25.2 |
|
|
Del(13q) |
37.0 |
55.8 |
<0.001 |
|
t(14;16) |
0.6 |
3.9 |
0.004 |
Chromosome 1q gain was identified as an independent variable for inferior overall survival (OS; hazard ratio [HR], 1.4; 95% confidence interval [CI], 1.097–1.787; p = 0.007), and progression-free survival (PFS; HR, 1.36; 95% CI, 1.16–1.58; p = 0.0003). No significant differences in PFS and OS were observed between patients with three copies of chromosome 1q (70.2%) and four or more copies of 1q (29.8%), with a p value > 0.05 for both.
Patients with high-risk mutations alongside 1q gain had shorter PFS (p = 0.0294) and OS (p = 0.0381). Presence of t(14;16), del(17p), or del(13q) with 1q gain reduced PFS and OS, while del(1p) alongside 1q gain was significant only for OS (Table 3).
Table 3. Risk association for patients with chromosome 1q gain alongside high-risk cytogenetic anomalies*
|
CI, confidence interval; del, deletion; HR, hazard ratio; OS, overall survival; PFS, progression-free survival T, translocation. |
||||
|
Concurrent cytogenetic anomaly |
OS |
PFS |
||
|---|---|---|---|---|
|
HR (95% CI) |
p value |
HR (95% CI) |
p value |
|
|
del(17p) |
1.5 (1.06─2.124) |
0.022 |
1.762 (1.191─2.607) |
0.005 |
|
t(14;16) |
3.277 (1.818─5.904) |
<0.001 |
3.288 (1.701─6.355) |
<0.001 |
|
del(13q) |
1.369 (1.088─1.723) |
0.007 |
1.385 (1.060─1.811) |
0.017 |
|
del(1p) |
1.547 (0.936─2.555) |
0.089 |
2.261 (1.317─3.882) |
0.003 |
From univariate and multivariate analysis, t(14;16), hypercalcemia, International Staging System (ISS) Stage III, and high lactate dehydrogenase were identified as risk factors for poor outcome in patients with chromosome 1q gain (Table 4). Describing patients with 1q gain as heterogeneous, Yang et al. established a risk-stratification score using these variables to classify patients with 1q gain into low, intermediate, and high-risk groups (low risk, 0 points; intermediate risk, 1–3 points; high risk, 4–7 points).
Table 4. Risk model for patients with chromosome 1q gain*
|
CI, confidence interval; HR, hazard ratio; ISS, International Staging System; LDH, lactate dehydrogenase; t, translocation. |
||||
|
Characteristic |
β value |
p value |
HR (95% CI) |
Points assigned† |
|---|---|---|---|---|
|
ISS III |
0.446 |
0.005 |
1.562 (1.141─2.138) |
1 |
|
Hypercalcemia |
0.803 |
<0.001 |
2.231 (1.530─3.254) |
2 |
|
Elevated LDH |
0.666 |
<0.001 |
1.946 (1.440─2.631) |
1 |
|
t(14;16) |
1.194 |
<0.001 |
3.302 (1.766─6.171) |
3 |
In the study, 376 patients were evaluable and stratified into low (31.6%), intermediate (61.7%), and high risk (6.7%) groups, with significantly different PFS (27.4, 18.1, and 6.1 months, respectively; p < 0.0001) and OS (59.8, 34.8, and 18.9 months, respectively; p < 0.0001), in association with early disease relapse. Differences were seen in terms of PFS and OS between the groups (Table 5).
Table 5. Relative hazard ratios of different risk criteria*
|
CI, confidence interval; HR, hazard ratio; OS, overall survival; PFS, progression-free survival. |
|||
|
Comparator |
HR |
95% Cl |
p value |
|---|---|---|---|
|
PFS |
|
|
|
|
Intermediate vs low |
1.74 |
1.32─2.29 |
<0.001 |
|
High vs intermediate |
2.78 |
1.77─4.37 |
<0.001 |
|
OS |
|
|
|
|
Intermediate vs low |
2.36 |
1.65─3.38 |
<0.001 |
|
High vs intermediate |
3.20 |
1.97─3.38 |
<0.001 |
There was a significantly lower rate of measurable residual disease (MRD) negativity in patients with chromosome 1q gain than in those without (41.9% vs 62.5%; p = 0.004). Achieving MRD negativity improved outcomes, including the:
- PFS (median, 35.1 vs 11.6 months; HR, 0.43; 95% CI, 0.32–0.58; p < 0.0001); and
- OS (median, not reached vs 25.4 months; HR, 0.45; 95% CI, 0.31–0.65; p < 0.0001) in patients with 1q gain.
Achieving MRD negativity also significantly improved outcomes, including:
- median PFS (31.4 vs 11.4 months; HR, 0.39; 95% CI, 0.22–0.68; p = 0.0008); and
- median OS (47.2 vs 23.6 months; HR, 0.31; 95% CI, 0.14–0.67; p = 0.0028) in intermediate/high-risk patients.
No differences were seen in PFS or OS in patients with or without 1q gain when MRD negativity was maintained.
Compared with those without 1q gain, median time-to-relapse of patients with 1q gain was significantly shorter (20.2 vs 28.3 months; p < 0.0001), with a significantly increased risk of relapse at both:
- 12 months (odds ratio, 1.87; 95% CI, 1.25–2.79; p = 0.002); and
- 24 months (odds Ratio, 1.93; 95% CI, 1.40–2.67; p < 0.001).
OS of patients with 1q gain was very strongly associated with relapse at 12 months (median, 14.2 vs 51.2 months; HR, 4.29; 95% CI, 3.16–5.82; p < 0.0001) and 24 months (median, 25.4 vs 67.0 months, HR, 5.09; 95% CI, 3.65–7.11; p < 0.0001).
No significant difference was seen in the objective response rate or the best response rate (at least a very good partial response) between patients with and without 1q gain after treatment with proteasome inhibitors and/or immunomodulatory drugs, or allogeneic hematopoietic stem-cell transplant.
Gain of chromosome 1q21 in patients with MM3
Kastritis et al.3 conducted a retrospective case note analysis of 912 patients treated at a single center between 2004 and 2021. At diagnosis, 249 (27.3%) patients had 1q21 gain, with three copies in 16.4% of patients and four or more copies in 10.9%. Patients with 1q21 gain had inferior PFS (median, 34 vs 20 months; p < 0.001) and inferior OS (median, 75 vs 44 months; p < 0.001).
The presence of 1q21 gain was associated with advanced ISS stage (p = 0.003) and concurrent presence of other cytogenetics abnormalities (p < 0.001). Multivariate analysis identified 1q21 gain as an independent prognostic factor for both PFS (p < 0.001) and OS (p = 0.008) (Table 6). The poorest OS was reported in revised ISS (R-ISS) Stage III patients with 1q21 gain compared with those without 1q21 gain (16 months vs 46 months; p < 0.001).
Table 6. Multivariate analysis of the effect of 1q21 gain adjusted for clinical variables *
|
ASCT, autologous stem cell transplant; CI, confidence interval; HDM, high-dose melphalan; HR, hazard ratio; IMiD, immunomodulatory drug; OS, overall survival; PI, proteosome inhibitor; PFS, progression-free survival; R-ISS, revised International Staging System. |
||||
|
Parameter |
β-level |
p value |
HR |
95% CI |
|---|---|---|---|---|
|
PFS |
||||
|
R-ISS III (reference value) |
— |
— |
1 |
— |
|
R-ISS I |
−0.873 |
<0.001 |
0.418 |
0.283─0.617 |
|
R-ISS II |
−0.578 |
0.001 |
0.561 |
0.404─0.779 |
|
Primary treatment |
|
|
|
|
|
HDM-ASCT |
−0.581 |
0.001 |
0.560 |
0.400─0.782 |
|
Induction: PI + IMiD |
|
0.037 |
|
|
|
Induction: PI +/− chemotherapy |
0.515 |
0.004 |
1.673 |
1.177─2.378 |
|
Induction: IMiD +/− chemotherapy |
0.464 |
0.012 |
1.591 |
1.109─2.284 |
|
1q21 gain present |
0.408 |
<0.001 |
1.503 |
1.207─1.872 |
|
OS |
||||
|
R-ISS III (reference value) |
— |
— |
1 |
— |
|
R-ISS I |
−1.210 |
<0.001 |
0.298 |
0.181─0.491 |
|
R-ISS II |
−0.551 |
0.005 |
0.576 |
0.394─0.884 |
|
Age <65 years |
−0.406 |
0.031 |
0.667 |
0.461─0.964 |
|
HDM-ASCT |
−0.653 |
0.003 |
0.521 |
0.340─0.796 |
|
Induction: PI +/− chemotherapy |
0.374 |
0.086 |
1.453 |
0.949─2.226 |
|
1q21 gain present |
0.344 |
0.008 |
1.410 |
1.092─1.821 |
Discussion
The study by Yang et al.2 validates 1q gain as an independent risk factor for poor prognosis in a large cohort of patients who received standard therapies (including proteosome inhibitors, immunomodulatory drugs, both, or hematopoietic stem cell transplantation). Gain of chromosome 1q demonstrates high heterogeneity in patients at diagnosis with MM, and can profoundly affect clinical outcomes, especially when 1q gain occurs concurrently with other cytogenetic anomalies, particularly t(14;16). A simple risk-scoring model based on the four easily identifiable variable, ISS III, hypercalcemia, high lactate dehydrogenase, and t(14;16), allows for the re-stratification of patients with NDMM with 1q gain.2 Further replications and validation of this risk tool in larger clinical trials could inform risk-adapted treatment for the subset of patients 1q gain.
Gain of 1q21 is associated with inferior PFS and OS and more advanced disease in patients with NDMM.3 Patients with R-ISS Stage III disease comprise an ultra-high-risk group with poor prognosis and may represent a subset of patients who require special consideration when selecting treatments.3
Similar data were identified by Tang et al.4, who identified reduced event-free survival (median, 35 vs 55 months; p = 0.05) and OS (median, 74 vs 168 months; p = 0.00025), in patients with 1q21 gain treated with bortezomib. Again, multivariate analysis established 1q21 gain as an independent risk factor for OS.4
Further study of chromosome 1 changes will not only affect OS and PFS in patients with NDMM through risk stratification, but also the development of novel therapeutic strategies.5 Analysis of chromosomal cytobands allows for the detection of changes in genes, such as BCL9, CKS1B, and ILF2, with potential utility as targets for novel agents (Figure 2).5
Figure 2. Analysis of chromosomal cytobands allows for the detection of changes in genes with potential utility as targets for novel agents*
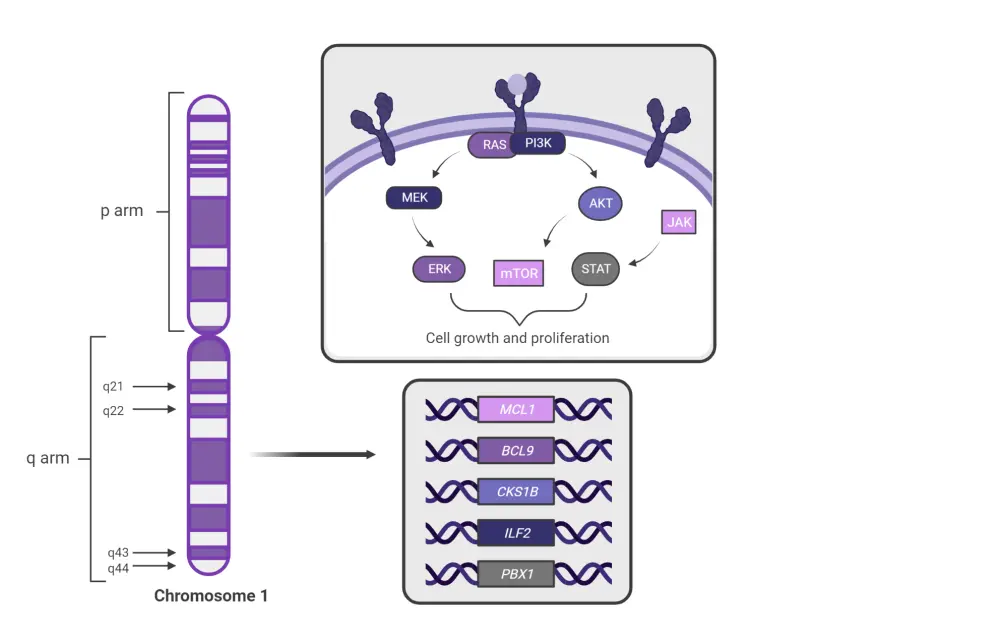
*Adapted from Sklavenitis-Pistofidis, et al.5 Created with BioRender.com.
Conclusion
Gain and loss of chromosome 1q has implications for patients with NDMM, affecting disease behavior, prognosis, and clinical outcomes according to the presence of other clinical features and cytogenetic abnormalities. Beyond current R-ISS guidelines for the risk stratification of patients with NDMM, consideration of other risk factors alongside the presence of highly heterogeneous chromosome anomalies will allow for the further stratification of high-risk and ultra-high-risk patient subsets. Chromosome 1q anomalies may also have a role to play in routine R-ISS risk stratification, which has recently been included in the second revision of the R-ISS guidelines, covered separately by the Multiple Myeloma Hub.
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content




